수원청개구리 (Hyla (Dryophytes) suweonensis Kuramoto)
|
|
▣ 국 명: 수원청개구리 ▣ 영 명: Suweon-tree frog ▣ 학 명: Hyla suweonensis ▣ 지정번호: 멸종위기야생동물 I급 ▣ 계 통: 양서강-무미목-청개구리과 |
1) 채집정보
- 채집허가: 2014.07.17 / 3마리
- 채 집 자: 서울대학교 민미숙
2) 채집현황 및 샘플처리
|
수원청개구리 채집현황 |
샘플처리 |
|||
|
채집일 |
개체 수 |
DNAseq |
RNAseq |
표본 |
|
2014.07.30 |
2 EA |
2 EA (몸통) |
2 EA (내장) |
- |
3) QC Data
|
샘플종 |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
수원청개구리 |
65.3 |
11.7 |
577.3 |
8.2 |
pass |
4) 기존 NCBI Taxonomy 유전정보 현황
|
유전정보 종류 |
Dryophytes 속 유전자원 |
Hyla suweonensis 유전자원 |
|
Nucleotide |
2,581 |
196 |
|
Protein |
1,508 |
150 |
|
Genome |
2 |
1 |
|
Gene |
50 |
37 |
5) Sequencing 분석 결과
- Illumina DNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
169,600,636 |
169,600,636 |
339,201,272 |
|
Number Of Bases |
|
|
21,369,680,136 |
21,369,680,136 |
42,739,360,272 |
|
error correction |
|||||
|
Number Of Reads |
|
|
161,523,082 |
159,313,522 |
320,836,604 |
|
Number Of Bases |
|
|
18,810,528,520 |
18,410,014,455 |
37,220,542,975 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
307,777,452 |
|
Number Of Bases |
|
|
|
|
35,940,123,938 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
13,059,152 |
|
Number Of Bases |
|
|
|
|
1,280,419,037 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
2,386,625 |
2,386,625 |
302,758 |
42,061 |
1,866 |
|
Number Of Bases |
668,192,717 |
668,192,717 |
228,066,795 |
54,836,871 |
4,476,961 |
|
Avg. Scaffold Size |
279 |
279 |
753 |
1,303 |
2,399 |
|
N50 Scaffold Size |
364 |
364 |
739 |
1,255 |
2,297 |
|
N80 Scaffold Size |
178 |
178 |
576 |
1,081 |
2,097 |
|
N90 Scaffold Size |
126 |
126 |
536 |
1,038 |
2,042 |
|
largest Scaffold Size |
5,914 |
5,914 |
5,914 |
5,914 |
5,914 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
63,439,777 |
3,643,748 |
93,857 |
3,389 |
44 |
|
Number Of Bases |
3,072,785,570 |
676,644,137 |
60,518,687 |
4,050,130 |
106,903 |
|
Avg. Contig Size |
48 |
185 |
644 |
1,195 |
2,429 |
|
N50 Contig Size |
42 |
193 |
622 |
1,155 |
2,223 |
|
N80 Contig Size |
33 |
124 |
538 |
1,046 |
2,061 |
|
N90 Contig Size |
32 |
110 |
518 |
1,020 |
2,013 |
|
Largest Contig Size |
5,328 |
5,328 |
5,328 |
5,328 |
5,328 |
- Illumina RNA Sequncing 결과
|
Input information |
|||||
|
|
Region1 |
Region2 |
Region3 |
Region4 |
Total |
|
raw data |
|||||
|
Number Of Reads |
116,457,440 |
116,457,440 |
9,406,614 |
9,406,614 |
251,728,108 |
|
Number Of Bases |
14,673,637,440 |
14,673,637,440 |
1,185,233,364 |
1,185,233,364 |
31,717,741,608 |
|
error correction |
|||||
|
Number Of Reads |
114,954,921 |
113,749,442 |
8,995,241 |
8,870,687 |
246,570,291 |
|
Number Of Bases |
14,272,528,339 |
14,087,354,291 |
1,101,410,323 |
1,082,508,754 |
30,543,801,707 |
|
pair reads |
|||||
|
Number Of Reads |
|
226,547,382 |
|
17,547,462 |
244,094,844 |
|
Number Of Bases |
|
28,122,222,192 |
|
2,152,123,166 |
30,274,345,358 |
|
single reads |
|||||
|
Number Of Reads |
|
2,156,981 |
|
318,466 |
2,475,447 |
|
Number Of Bases |
|
237,660,438 |
|
31,795,911 |
269,456,349 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
133,584 |
133,584 |
29,564 |
13,534 |
4,633 |
|
Number Of Bases |
60,692,315 |
60,692,315 |
38,761,572 |
27,592,745 |
15,254,131 |
|
Avg. Scaffold Size |
454 |
454 |
1,311 |
2,038 |
3,292 |
|
N50 Scaffold Size |
847 |
847 |
1,583 |
2,173 |
3,255 |
|
N80 Scaffold Size |
256 |
256 |
816 |
1,361 |
2,399 |
|
N90 Scaffold Size |
166 |
166 |
640 |
1,172 |
2,181 |
|
largest Scaffold Size |
28,871 |
28,871 |
28,871 |
28,871 |
28,871 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
1,836,259 |
185,405 |
28,561 |
10,203 |
2,408 |
|
Number Of Bases |
163,179,459 |
62,590,624 |
30,190,736 |
17,524,641 |
6,919,325 |
|
Avg. Contig Size |
88 |
337 |
1,057 |
1,717 |
2,873 |
|
N50 Contig Size |
91 |
471 |
1,157 |
1,733 |
2,773 |
|
N80 Contig Size |
50 |
190 |
691 |
1,230 |
2,241 |
|
N90 Contig Size |
47 |
147 |
586 |
1,110 |
2,114 |
|
Largest Contig Size |
11,703 |
11,703 |
11,703 |
11,703 |
11,703 |
(2) 유전체 서열확보 : 현재 20종 확보
|
Sample |
구분 |
raw data size |
trim data size |
비고 |
|
수원청개구리 |
DNA |
42,739,360,272 |
37,220,542,975 |
|
|
RNA |
31,717,741,608 |
30,543,801,707 |
|
(3) 기타분석
A. Genome Size estimation
|
샘 플 명 |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
수원청개구리 |
23 |
26,457,699,216 |
190 |
139,251,048 |
243.59 |
B. 반복서열
- 시퀀싱이 완료된 수원청개구리 DNA 및 RNA의 contigs 중 1kb 이상의 contigs의 반복서열을 탐색(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 3,389 |
|
sequences: 9,723 |
||||||||
|
total length: 4,050,130 bp (4,050,130 bp excl N/X-runs) |
|
total length: 16,747,459 bp (16,747,459 bp excl N/X-runs) |
||||||||
|
GClevel: 41.73% |
|
GC level: 46.44% |
||||||||
|
bases masked: 7,336bp ( 0.18%) |
|
bases masked: 9,020 bp ( 0.05%) |
||||||||
|
|
numberof elements |
length occupied |
percentage of sequence |
|
|
numberof elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
4 |
515 bp |
0.01% |
|
SINEs: |
3 |
138 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
0 |
0 bp |
0.00% |
|
|
MIRs |
1 |
35 bp |
0.00% |
|
LINEs: |
27 |
3,080 bp |
0.08% |
|
LINEs: |
26 |
2,953 bp |
0.02% |
||
|
|
LINE1 |
19 |
2,121 bp |
0.05% |
|
|
LINE1 |
9 |
1,251 bp |
0.01% |
|
|
LINE2 |
3 |
564 bp |
0.01% |
|
|
LINE2 |
4 |
900 bp |
0.01% |
|
|
L3/CR1 |
4 |
334 bp |
0.01% |
|
|
L3/CR1 |
8 |
565 bp |
0.00% |
|
LTRelements: |
7 |
1,469 bp |
0.04% |
|
LTRelements: |
13 |
1,784 bp |
0.01% |
||
|
|
ERVL |
1 |
557 bp |
0.01% |
|
|
ERVL |
1 |
83 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
1 |
52 bp |
0.00% |
|
|
ERV_classI |
4 |
527 bp |
0.01% |
|
|
ERV_classI |
9 |
686 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
DNAelements: |
13 |
1,638 bp |
0.04% |
|
DNAelements: |
18 |
4,091 bp |
0.02% |
||
|
|
hAT-Charlie |
3 |
670 bp |
0.02% |
|
|
hAT-Charlie |
3 |
1,421 bp |
0.01% |
|
|
TcMar-Tigger |
6 |
446 bp |
0.01% |
|
|
TcMar-Tigger |
6 |
1,280 bp |
0.01% |
|
Unclassified: |
1 |
40 bp |
0.00% |
|
Unclassified: |
0 |
0 bp |
0.00% |
||
|
Total interspersed repeats: |
|
6,742 bp |
0.17% |
|
Total interspersed repeats: |
|
8,966 bp |
0.05% |
||
|
Small RNA: |
8 |
839 bp |
0.02% |
|
Small RNA: |
2 |
120 bp |
0.00% |
||
|
Satellites: |
2 |
207 bp |
0.01% |
|
Satellites: |
0 |
0 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
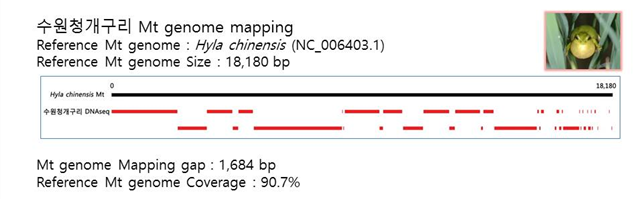
D. Mito-genome Map
(4) Microsatellite 후보군
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
수원청개구리 |
185 |
49 |
7 |
- |
- |
- |
- |
- |
- |
241 |
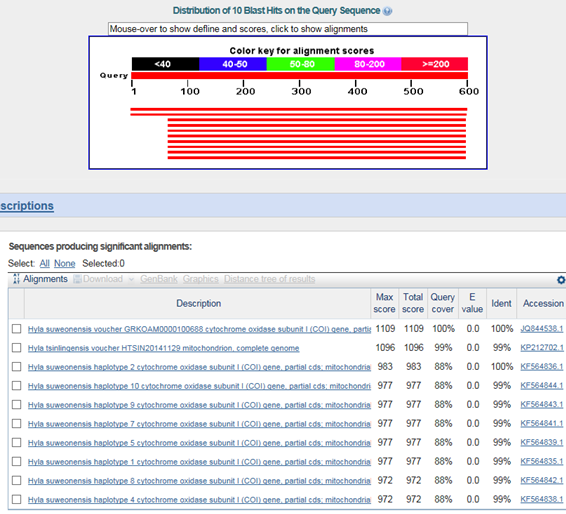
B. 바코드서열
>수원청개구리(Hyla suweonensis)
GGGATAGTTGGCACTGCCCTCAGCCTCCTAATTCGAGCAGAACTTAGCCAGCCTGGTTCC
CTTCTGGGTGATGACCAGATCTACAATGTTATCGTAACAGCCCATGCCTTTGTTATAATT
TTCTTCATAGTAATACCAATTCTAATCGGGGGCTTCGGCAACTGACTAGTTCCTTTAATA
ATCGGGGCCCCTGATATAGCCTTCCCACGAATAAACAATATAAGCTTTTGGCTTCTTCCA
CCATCTTTTCTTCTTCTTCTAGCGTCAGCCGGAGTCGAAGCAGGAGCTGGAACAGGATGA
ACCGTCTACCCCCCGCTTGCCGGAAATCTGGCCCACGCCGGACCATCCGTTGACTTAACT
ATTTTCTCTCTTCACCTAGCAGGGGTTTCTTCGATTTTAGGCGCCATTAATTTTATCACC
ACAATCCTCAATATAAAACCCCCGTCAATAACACAATACCAAACACCCCTCTTTGTGTGA
TCCGTTCTCATTACTGCCGTACTGCTTCTCTTATCCCTTCCAGTCTTAGCAGCAGGAATC
ACTATACTGCTCACTGATCGAAACTTAAACACAACATTCTTTGACCCAGCAGGAGGAGGA